Visualize the scatter of minimization results – prof-plotresultscatter¶After the goodness of fit function was minimized with prof-tune, prof-plotresultscatter can be used to visualize the scatter of the minimization results obtained with different sets of anchor points. This is necessary to check that the minimizer does not get caught in local minima and that the goodness of fit (GoF) does not depend strongly on the combination of anchor points/MC runs. The plots produced show the scatter of the results in the GoF-parameter plane. For each parameter one plot is produced. For a good tuning one expects that the minimization results cluster around the minimization result that was obtained with all MC runs as anchor points. If more than one cluster appears professor can be used interactively to extract the cluster around the all-MC-runs result by imposing simple cuts on the parameter values of the minimization results (see: professor.minimize.result.ResultList.filtered()). Possible reasons for a wide spread of the minimization results in one ore more parameters are
The original sampling ranges can be plotted using --ranges RANGEFILE to check that the results are not in regions of extrapolation. The styles of the points changes by default with the number of anchor points used for the polynomial approximations, e.g. results obtained from 10 anchor points are displayed in a different style than results from 20 anchor points. A second criterion that changes the display style is if parameters were limitted during the minimization or not. A third, optional criterion is to discriminate the results by the method that was used to assign the initial point for the minimization, e.g. center or random. The location for the output files can be chosen with --outdir DIRECTORY. For each parameter that was tuned one file is created. The file names are constructed from the parameter names. In addition --suffix can be used to add an individual tag to the file names. The results that are plotted are taken from the result files that are given as arguments to the scripts. If multiple result files are given, the results from these files are concatenated. By default PDF files are created using matplotlib. --make-plots can be used to produce dat files for make-plots instead. Example¶Simply plot the scatter of results from myresults.pkl and draw lines at the parameter boundaries in sample.ranges: prof-plotresultscatter --ranges sample.ranges myresults.pkl
Command-line options¶The --datadir DATADIR and related options are used as normal to specify the reference data, MC runs, and interpolation objects: see the path options page.
Display style changes¶
|
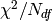 (default) or
(default) or 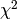 as GoF.
as GoF.